Single-cell RNA sequencing, or scRNA-seq, has revolutionized the way scientists study cells, by grouping them based on similar gene expression. Although several scRNA-seq studies have explored bone physiology, they often lacked sufficient data on some cell populations and reached divergent conclusions, assigning different names to similar cell clusters. These discrepancies hinder research efforts and complicate the understanding and treatment of bone-related diseases.
Intawat Nookaew and Charles O’Brien, both professors at the University of Arkansas for Medical Sciences, have studied bone pathophysiology for years and recently collaborated to analyze scRNA-seq results from bone, integrating their own data with that of other researchers. Their results were published in the Journal of Biological Chemistry.
Previous scRNA-seq studies of bone were unable to obtain large numbers of osteoblasts, the cells responsible for building new bone, and osteocytes, the cells derived from osteoblasts that are involved in controlling bone remodeling.
“We had two main goals with this paper,” O’Brien said. “We wanted to increase the number of osteoblasts and osteocytes in single-cell RNA sequencing data, and the second was to put our results together with the results of five or six other big studies and try to normalize them together to find out how many different cells types we’re talking about and come up with a consistent nomenclature for them.”
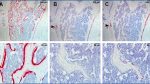
Nookaew et al./JBC – This image from the paper shows six types of bone cells identified by the authors from a mouse femoral bone at lower and higher magnifications.
O’Brien and Nookaew focused on increasing osteoblast and osteocyte scRNA-seq data by collecting cells from the femurs and tibias from two mouse strains, Osx1-Cre and Dmp1-Cre, which are popularly used to study osteoblasts and osteocytes, respectively.
After the cells were sequenced, O’Brien and Nookaew combined their scRNA-seq data with other studies and condensed them, boiling about 20-30 cell types down to 11 major cell clusters.
“There was one population in particular that seemed to be the same cell type, but each paper gave it a different designation, which could lead to difficulties in the future because there’s not a consistent nomenclature,” O’Brien said.
Their efforts mark an initial stride toward standardizing cell nomenclature within skeletal research and medicine.
“That is why we tried to harmonize a lot of data sets to show that these studies are comparable,” Nookaew said.
One major challenge was extracting enough osteocytes from mouse femurs and tibias, Nookaew said, explaining, “Osteocytes are in the bone matrix and are difficult to get out. Previous studies didn’t have a lot of osteocytes, and that’s why in our study we generated about 2,000 osteocytes so that others can see and use the data.”
They developed a cell map that can be used as a starting point for others who are studying diseases affecting the skeletal system.
Osteoblast production and efficiency decline with age, leading to loss of bone mass, but researchers don’t yet understand how this happens. O’Brien and Nookaew believe they can unlock the answers by using single-cell RNA-seq.
“By taking the information we’ve learned, we’re using it to study different populations of cells that may become osteoblasts and osteocytes and how those populations change with age,” O’Brien said.
Nookaew et al./JBC – This image from the paper shows vertebral bone sections from mice treated with vehicle, top, or denosumab, bottom.
Nookaew and O’Brien used the Osx1-Cre and Dmp1-Cre mouse strains which are genetically modified to study osteoblasts and osteocytes; however, they found that these strains are not as specific to osteoblasts and osteocytes as previously described, showing that many other cells in the bone can be targeted.
“This is a very significant and, until recently, not very well recognized limitation of any Cre technology,” O’Brien said. “You have to be careful about the conclusions that you draw, as you’ve eliminated or activated a gene of interest in many other cell types in addition to osteoblasts and osteocytes.”
Moving forward, Nookaew and O’Brien have identified genes that are relatively specific to different cell types and are now developing new Cre mouse strains that will allow skeletal researchers to genetically target a subset of cells.
Source – American Society for Biochemistry and Molecular Biology
